Researcher asks what makes a species by looking closely at lizards

LAWRENCE — With a recent four-year, $600,000 grant from the National Science Foundation, a University of Kansas researcher is undertaking the most-detailed analysis ever carried out of how reptiles branch into various species.
In doing so, Richard Glor, associate professor of ecology and evolutionary biology at KU and associate curator of herpetology at KU’s Biodiversity Institute, is questioning some of the very methods scientists use to define species.
Further, by combining laboratory studies with fieldwork and genomic sequence data, Glor and his team hope to determine the genetic basis for species differences and why these species diverged in the first place.
Glor said biologists traditionally employ the idea of “reproductive isolation,” or the inability of one kind of animal to fruitfully breed with another, to determine boundaries defining individual species.
“Every student in biology has learned the philosophy of what a species is — something that can’t reproduce with other species,” Glor said. “So when someone describes a new species, you’d think they’ve done a bunch of crossing experiments [to see if a species can reproduce with another], but the answer is that nobody ever does that.”
He said more laboratory studies should be conducted to verify assumptions about boundaries between species and less reliance should rest on how animals look to the eye. His new grant is based on the idea that conventional methods have overlooked recent and ongoing speciation events.
“Traditional taxonomists say that if species exhibit differences in their appearance — this one is green, and this one is red — they likely aren't reproducing with one another,” Glor said. “I saw that as an understudied area in speciation research. We always talk about reproductive isolation, but we never test it.”
Glor has revealed this shortcoming in part by conducting crossing experiments on a group of lizards called bark anoles found on Hispaniola, the Caribbean island shared by Haiti and the Dominican Republic. These lizards are notable for the remarkable variation of their “dewlap,” or flap of skin along the throat, that can be found in different colors and patterns.
“The dewlap is colorful, kind of like a fancy car or jewelry in the human population,” Glor said. “It shows the female you’re attractive and helps her recognize her own species, so it’s critical to sexual selection. The group that I study is particularly noteworthy because a single species exhibits very different dewlap colors and patterns (i.e., some populations are yellow, while others are red). Most anole species exhibit only a single color.”
Previous biologists have relied on variation in dewlaps to separate bark anoles into different subspecies.
“When herpetologists drove across Haiti's mountainous Tiburon Peninsula more than 50 years ago, they saw bark anoles with pale yellow dewlaps at the beginning of their journey, bark anoles with orange and then wine red dewlaps further along, and then started seeing bark anoles with pale yellow dewlaps again by the time they reached the far tip of the peninsula,” Glor said. “As a result, they described these populations with different dewlap colors as distinct subspecies.”
But as Glor and his colleagues obtained genomic data from the lizards, they found dewlap colors weren’t a trustworthy sign of the underlying genetic differences scientists expect to find in distinct species.
“These populations don’t seem to be genomically differentiated,” he said. “Some of those taxa that traditional methods show to be different species probably aren’t different species. Natural selection is driving them to have different dewlap color and appearance for reasons not related to them being different species.”
In other cases, Glor said traditional methods missed species that appear to be strongly differentiated genetically because they are nearly impossible to distinguish externally.
Glor’s work represents the most detailed and integrated analysis ever performed on squamate reptiles — a group of almost 10,000 species, including all lizards and snakes. The research will result in the establishment of a new laboratory model organism, the first squamate genome assembled through linkage mapping and the first “estimates of heritability and the genomic basis for phenotypic traits” vital to systematically studying reptiles in general.
Further, the research has resulted in a public exhibit now on display at the KU Museum of Natural History.
Ultimately, however, Glor said that evolution in species is an unending and fluid process that biologists might always struggle to catalogue and measure.
“All of these ideas like reproductive isolation or morphological divergence are all just benchmarks that occur as new species are forming,” he said. “It could be that some species exhibit all these benchmarks, and some display only one or two. The problem identifying species is that we're trying to put discrete start and stop points on a process we know is continuous.”
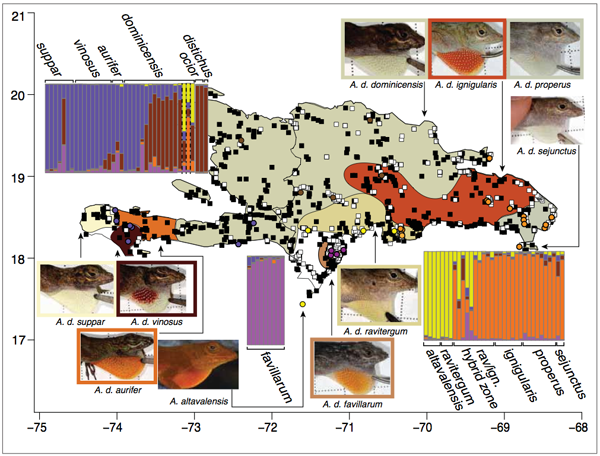
Graphic: Taxonomic distributions, sampling, and preliminary results. Images illustrate dewlap coloration typical of each of the Hispaniolan taxa assigned to the distichus species group. Shaded regions of the map indicate ranges of subspecies of A. distichus and correspond with the prevailing color of the dewlap in each of these subspecies. Filled black squares indicate sampling localities from which whole animal voucher specimens are available. White squares indicate localities from which tissue samples, whole animal voucher specimens and dewlap photographs are available. Bar graphs were generated by analyses of AFLP loci obtained from 96 individuals sampled from across the range of A. distichus in STRUCTURE and indicate proportion genotypic assignment of each individual to one of five genotypic clusters. Colored circles indicate localities from which AFLP data were obtained, with the color of each circle corresponding with the majority genotype assignment for individuals at a given locality.
Photo: KU is celebrating 100 years of herpetology research. World-renowned herpetologist Rich Glor is a curator of the collection. Credit: Meg Kumin, KU Marketing Communications.