Study finds 'black box' methods used by biologists probably overestimate number of new species

LAWRENCE — Do researchers need to “hit the brakes a little” before declaring a species is new to science?
A recent University of Kansas study published in the peer-reviewed journal Molecular Ecology demonstrates the misuse and abuse of methods scientists commonly use to place boundaries between different species, especially so-called “cryptic” species, could lead to the overestimation of species diversity.
KU researchers assessed genetic differences between populations of Malaysian torrent frogs using two approaches: First, they employed species delimitation methods (SDMs) that focus on identification and demarcation of genetic breaks in targeted gene regions (DNA sequences) to determine species boundaries. Second, they used data gathered from across the genome to perform a comprehensive analysis of molecular variation — to determine whether targeted gene studies reflect real “species.”
The new study, performed by doctoral student Chan Kin Onn and his adviser, KU Professor Rafe Brown, found that SDMs inferred up to seven separate species, while the more exhaustive analysis instead suggested the statistically justifiable existence of just three species, including various populations of the same frog species, which had a history of trading genes back and forth.
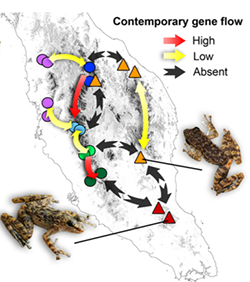 “SDMs were effective at delimiting divergent lineages in the absence of gene flow but overestimated species in the presence of marked population structure and gene flow,” the authors wrote. Gene flow is the exchange of genes from one population to another — such as exchange of genetic material among frogs living on different mountain ranges, facilitated by migration of individuals among populations.
“SDMs were effective at delimiting divergent lineages in the absence of gene flow but overestimated species in the presence of marked population structure and gene flow,” the authors wrote. Gene flow is the exchange of genes from one population to another — such as exchange of genetic material among frogs living on different mountain ranges, facilitated by migration of individuals among populations.
“We’re not calling into question analyses or methods of the past but, rather, we espouse a more cautious implementation and interpretation,” said Chan Kin Onn, of KU’s Biodiversity Institute and Department of Ecology & Evolutionary Biology, who led the new study. “We’re not saying all those papers describing new species are wrong. I’ve described species using SDMs myself. But we suggest that, in some cases, we should be more cautious when describing new species, especially when gene flow among species is involved. Most of the available programs do not account for gene flow, so if genetic exchange is present, it’s a no-brainer. We could easily get erroneous results. We suggest in this study that success is likely a matter of using the right tool for the right task.”
Chan and Brown’s co-authors were L. Lee Grismer from La Sierra University, Yong-Chao Su of Kaohsiung Medical University in Taiwan, Jesse Grismer of Auburn University and UCLA, Evan Quah of Universiti Sains Malaysia and Alana Alexander of KU.
The researchers performed their comparative analyses on populations of a group of Malaysian torrent frogs of the genus Amolops, all of which were initially believed to be of the same species.
“These torrent frogs are widely distributed across most of Asia,” Chan said. “Previous studies treated them as 51 species. However, in Peninsular Malaysia, past analyses suggested there was only one species. Our study started when we surveyed the peninsula using methodology from our usual surveys — we found these frogs in abundance in clean forest streams. They were so common we didn’t pay them a lot of attention. Over time, however, we noticed that different populations looked strikingly physically different. We wondered if there could be more than one species in Malaysia.”
Chan decided to examine the torrent frogs using both commonly used SDMs and a more modern populations genomics approach to see if there was a difference between results.
“This was a methodological comparison study,” Chan said. “I wanted to examine the efficacy of traditional approaches for identifying new species. I was motivated to do so because ‘new species’ are being described like crazy in Asia — but most of these studies look at the same kind of data, use the same kind of analysis and employ an almost formulaic template in ‘species delimitation’ studies. However, most of these species delimitation approaches don’t account for gene flow between populations — a real, biological phenomenon. Because of this, species delimitation models may not properly fit studies of divergence when gene flow is present."
The team analyzed 225 torrent-frog samples using morphological, mitochondrial DNA and genome-wide single-nucleotide polymorphism data to objectively find species boundaries while accounting for past and present gene flow between the frog populations.
“In today’s age of genomic science, it’s really important to look at genetic structure in finer detail,” Chan said. “It’s no longer sufficient to merely identify genetic ‘breaks’ between populations to justify their recognition as ‘new species.’ We also need to examine how genes are moving among populations and how that affects the integrity of species boundaries.”
Next, Chan will use the findings from Peninsular Malaysian torrent frogs as the basis to conduct a larger study on Malaysian karst geckos. He recently received a $19,500 grant from the National Science Foundation to test the hypothesis of ecological speciation in bent-toed geckos.
“The NSF Dissertation Improvement Grant program made my dissertation possible,” Chan said. “It’s a shame to learn the NSF discontinued the program this year. How will future U.S. scientists learn how to write NSF grant proposals?”
Chan said his work on peninsular Malaysian torrent frogs was a “proof-of-concept” for his larger dissertation study with the geckos.
“I wanted to use genomic methods to see if they were able to accurately delineate cryptic species boundaries as a test run,” Chan said. “It was hugely successful for a system of torrent frogs, which was thought to include only one species. Now, we’re going to apply the same framework to a vastly different system, including those that potentially could be overdescribed.”
Chan’s collaborative work with Lee Grismer of La Sierra University, spearheading research on geckos of Malaysia (Chan’s home country), has helped to prioritize conservation efforts and even influenced landscape use and management plans related to quarrying karst formations in order to preserve a new, threatened species’ habitat.
“One of the main reasons we’re doing this is because we’ve found so many species on isolated karst formations, whose habitats are threatened by being ground up for cement,” Chan said. “We have discovered and described new species that are already known to be under risk of extinction. This is because the only available habitat is being quarried for massive commercial cement production. We managed to halt one of these projects a few years ago. Some conservation organizations contacted us, saying, ‘You guys have to come and survey our mountain — the cement companies want to grind it down to dust.’ We visited the sites and found new species of lizards found nowhere else on Earth — and the cement company backed out.”
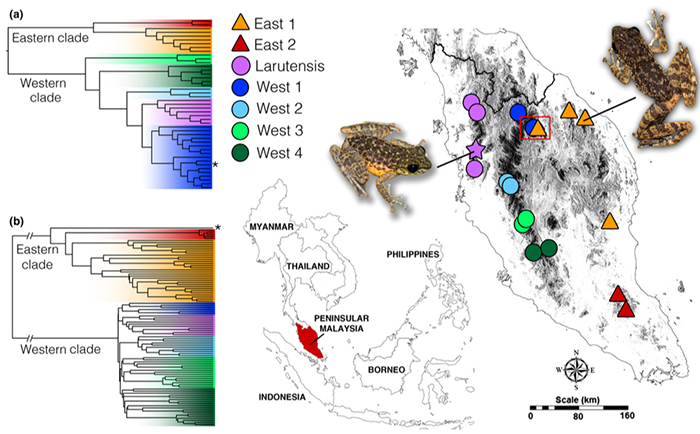
At top: photo illustration by Brendan Lynch, KU News Service.
Top right: Contemporary gene flow scenarios based on plausible phylogenetic relationships, population structure and geography. Source: Chan Kin Onn
Bottom: Likely evolutionary diversification based on analysis of rRNA-encoding mitochondrial gene. Source: Chan Kin Onn