Using fungi, researchers convert ocean plastic into ingredients for drug industry
LAWRENCE — Research on fungi underway at the University of Kansas has helped transform tough-to-recycle plastic waste from the Pacific Ocean into key components for making pharmaceuticals.
The chemical–biological approach for converting polyethylene uses an everyday soil fungus called Aspergillus nidulans that has been genetically altered. The results were reported recently in the paper “Conversion of Polyethylenes into Fungal Secondary Metabolites” published in Angewandte Chemie, a journal of the German Chemical Society.
“What we've done in this paper is to first digest polyethylenes using oxygen and some metal catalysts — things that are not particularly harmful or expensive — and this breaks the plastics into diacids,” said co-author Berl Oakley, Irving S. Johnson Distinguished Professor of Molecular Biology at KU.
Next, long chains of carbon atoms resulting from the decomposed plastics were fed to genetically modified Aspergillus fungi. The fungi, as designed, metabolized them into an array of pharmacologically active compounds, including commercially viable yields of asperbenzaldehyde, citreoviridin and mutilin.
Unlike previous approaches, Oakley said the fungi digested the plastic products quickly, like “fast food.”
“The thing that’s different about this approach is it's two things — it's chemical, and it's fungal,” he said. “But it's also relatively fast. With a lot of these attempts, the fungus can digest the material, but it takes months because the plastics are so hard to break down. But this breaks the plastics down fast. Within a week you can have the final product.”
The KU researcher added the new approach was “bizarrely” efficient.

“Of the mass of diacids that goes into the culture, 42% comes back as the final compound,” he said. “If our technique was a car, it would be doing 200 miles per hour, getting 60 miles per gallon, and would run on reclaimed cooking oil.”
Previously, Oakley has worked with corresponding author Clay Wang of the University of Southern California to produce about a hundred secondary metabolites of fungi for a variety of purposes.
“It turns out that fungi make a lot of chemical compounds, and they are useful to the fungus in that they inhibit the growth of other organisms — penicillin is the canonical example,” Oakley said. “These compounds aren’t required for the growth of the organism, but they help either protect it from, or compete with, other organisms.”
For a time, scientists thought they’d fully exploited the potential of fungi to produce these compounds. But Oakley said the age of genome sequencing has unlocked new possibilities for using secondary metabolites to benefit humanity and the environment.
“There was a realization there were lots and lots of clusters of genes that made secondary metabolites that nobody had discovered — and there are millions of species of fungi,” Oakley said. “A lot of companies have done good work over the years, but it was very much incomplete, because they were just growing things in the incubator and examining them for production of new compounds — but 95 percent of the gene clusters were just silent since they are not ‘turned on’ until needed. They weren't doing anything. So, there are lots more things to discover.”
Oakley’s lab at KU has honed gene-targeting procedures to change the expression of genes in Aspergillus nidulans and other fungi, producing new compounds.
“We’ve sequenced the genomes of a bunch of fungi now, and we can recognize the signatures of groups of genes that make chemical compounds,” he said. “We can change the expression of genes; we can remove them from the genome; we can do all kinds of things to them. We could see there were lots of these secondary metabolite gene clusters there and our gene-targeting procedures allowed us, at least in principle, to turn some of those clusters on.”
Oakley and Wang’s co-authors were Chris Rabot, Yuhao Chen, Swati Bijlani, Yi-Ming Chiang and Travis Williams of USC, and Elizabeth Oakley of KU.
The researchers focused on developing secondary metabolites to digest polyethylene plastics because those plastics are so hard to recycle. For this project, they harvested polyethylenes from the Pacific Ocean that had collected in Catalina Harbor on Santa Catalina Island, California.
“There’ve been a lot of attempts to recycle plastic, and some of it is recycled,” Oakley said. “A lot of it is basically melted and spun into fabric and goes into various other plastic things. Polyethylenes are not recycled so much, even though they're a major plastic.”
The KU investigator said the long-term goal of the research is to develop procedures to break down all plastics into products that can be used as food by fungi, eliminating the need to sort them during recycling. He added the work is emblematic of KU’s Earth, Energy + Environment research theme, geared toward “increasing understanding to help sustain the life of our planet and its inhabitants.”
“I think everybody knows that plastics are a problem,” Oakley said. “They’re accumulating in our environment. There’s a big area in the North Pacific where they tend to accumulate. But also you see plastic bags blowing around — they’re in the rivers and stuck in the trees. The squirrels around my house have even learned to line their nest with plastic bags. One thing that's needed is to somehow get rid of the plastic economically, and if one can make something useful from it at a reasonable price, then that makes it more economically viable.”
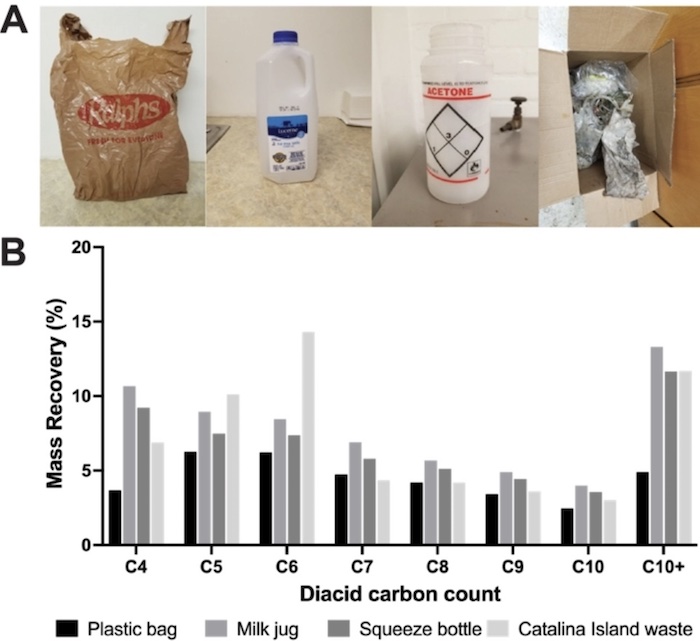
Top photo: Post-consumer plastics degraded in this study. A) From left to right: LDPE plastic grocery bag, HDPE milk jug, LDPE laboratory squeeze bottle, Pacific gyre waste collected from Santa Catalina Island, California. B) The distribution of diacid products after post-consumer polyethylene waste degradation using the optimized reaction. Credit: Rabot, et al.